Company Insight
Sponsored by Syngene
Direct to Biology for Drug Discovery Acceleration: Syngene’s D2B Platform
Braja Gopal Das, Subhendu K. Mohanty and Chantal Alcouffe
Main image: Biosecure Act 2025 and its implications for biopharma supply chain risk and regulatory compliance. Generated by an LLM based on given prompts
The BIOSECURE Act is now law, folded into the FY26 Senate NDAA on December 18, 2025. The Biosecure Act will phase in government-wide restrictions on the use of “biotechnology equipment or services’’ from designated biotechnology companies of concern, affecting federal contracts, grants, and loans, and creating new biopharma supply chain risk.
Primarily, the BIOSECURE Act introduces two core restrictions for US executive agencies. First, agencies must not procure or obtain biotechnology equipment or services from a designated biotechnology company of concern. Second, agencies must not enter, extend, or renew a federal contract with an entity that uses covered biotechnology equipment or services in performing the federal contract. The enacted language also extends restrictions to certain federal loan and grant funds used to procure covered items or to contract in a way that would require the use of those items.
A key point is timing. The restrictions will roll out in stages. First, the government is expected to publish a list of designated companies, then issue guidance and update federal procurement rules. The restrictions will start after set lead times. Some existing contracts may be allowed to continue for a limited period, but they will still face transition deadlines later.
Why drug discovery acceleration needs faster cycles
In drug discovery, speed matters, but speed alone is not enough. Teams also need reliable data early, so they can decide which chemical series deserve more effort and which ones should be stopped. Traditional drug discovery has served the industry well, but it is often slowed by one practical issue: compounds usually need to be synthesized, purified, characterized, and only then tested in biology. That sequence creates a real bottleneck. Direct to biology, or D2B, has gained attention because it challenges that old rhythm and supports drug discovery acceleration. Instead of waiting for full purification, crude reaction mixtures are screened directly in suitable biological assays, helping teams move from synthesis to insight much faster and improving the DMTA (Design, Make, Test and Analyse) cycle in early discovery. This approach is a way to accelerate hit identification and make faster decisions by removing a major bottleneck in the discovery pipeline.
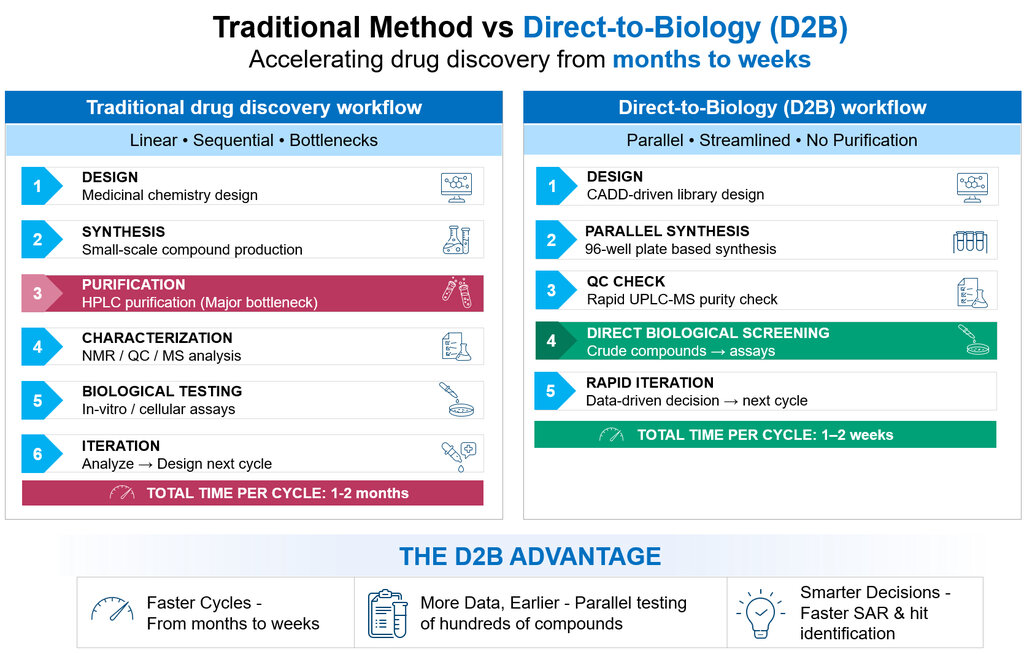
Comparison of traditional hit-to-lead and direct-to-biology workflows
What makes direct to biology different
The real difference between D2B and a traditional medicinal chemistry workflow is not just the use of crude material. It is the mindset behind the workflow. Traditional medicinal chemistry work is usually built around making a smaller number of purified compounds with high analytical confidence before biological testing. D2B flips that order for early discovery. It asks a practical question: can we get a biologically meaningful answer earlier, even before purification, and use that answer to guide the next round? When the chemistry is well chosen, and the assay is carefully designed, the answer is often yes. Moreover, D2B has developed a plate-based synthesis approach, enabling the generation of a vast number of compounds in a single run by exploring all possible variables from minimal starting material. This method significantly reduces both cost and time compared to traditional batch synthesis for the same number of compounds. D2B is now being used to explore chemical space more broadly, reduce cycle times, and support a faster DMTA cycle. At the same time, the best D2B platforms do not ignore quality; they use smart QC and data interpretation to keep speed from becoming noise.
Syngene’s integrated direct to biology platform
Syngene’s D2B capability is built as an integrated platform rather than a single high-throughput chemistry step. The base material shows a connected workflow that starts with software-driven compound design, moves into parallel microscale synthesis in 96-well plates, uses automated UPLC-MS for quality control, and then feeds compounds into biological and DMPK assays. That matters because in real projects, speed comes from handoffs being smooth, not from one fast experiment in isolation. Syngene’s CADD workflow adds another layer of value by using cheminformatics library enumeration, and property-based filtering to select compounds worth making in the first place. In simple terms, this means the platform is not only fast at making compounds; it is also thoughtful about which compounds enter the cycle.
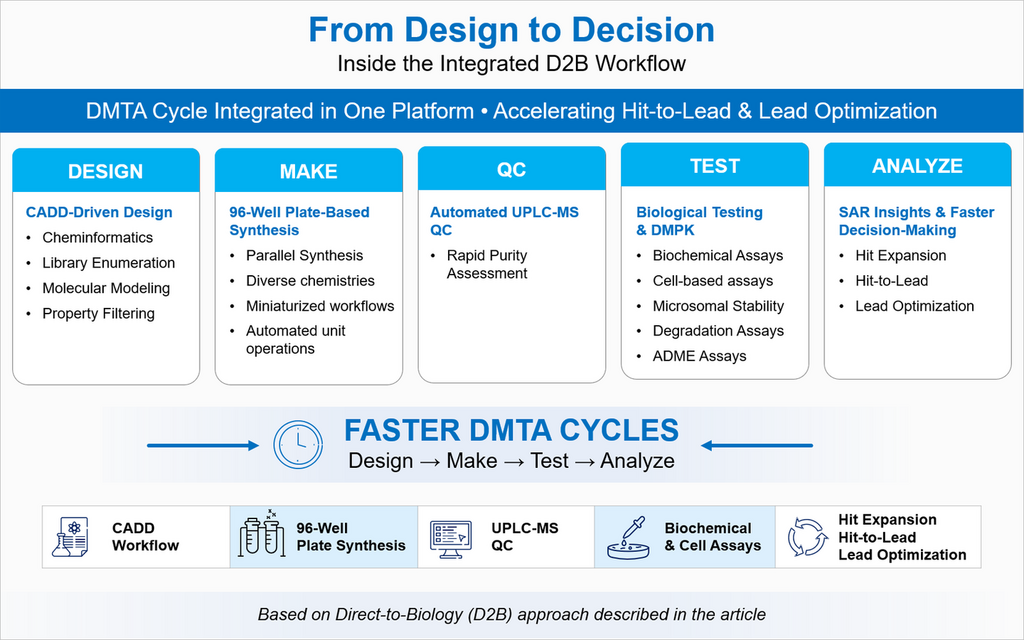
Integrated Direct-to-Biology workflow showing CADD-driven design, parallel plate-based synthesis, UPLC-MS quality control, and biochemical and cell-based assays to support faster DMTA cycles.
Where Syngene has a clear advantage in direct to biology
Syngene’s platform becomes especially powerful in targeted protein degradation work. The attached material highlights integration with Syngene’s internal SYNTAC library of E3 binders and linkers, which supports rapid PROTAC design and SAR exploration. This fits well with the wider direction of the field, where such modular systems benefit from rapid screening and iteration. In one example from the base material, Syngene prepared 48 PROTAC degraders through a one-step amide coupling campaign in just a few days, and 10 showed strong degradation potency. That is exactly the type of output clients look for: faster progression, broader exploration, and earlier identification of promising matter.
To learn more and register for the mAb accelerator program, click here: mAb Accelerator Program - Syngene International Ltd
Beyond simple one-step chemistry in hit to lead optimization
A common criticism of D2B is that it works only for very simple chemistry. Syngene’s material addresses that directly. It describes the successful application of the platform to a four-step sequence that delivered a 24-member library, with three compounds showing meaningful target degradation activity. That is important because it shows practical flexibility. The chemistry menu listed for the platform, including deprotections, sequential amide couplings, reductive amination, and urea synthesis, also suggests that the system is designed for real medicinal chemistry problems rather than narrow demonstrations. This wider reaction compatibility strengthens the case that Syngene can use D2B not just for quick screening exercises, but as a serious engine for hit expansion and hit to lead optimization.
Why this matters for pharma and biotech in drug discovery acceleration
The market is moving toward faster, more integrated discovery support. Pharmaceutical and biotech companies are increasingly looking for partners who can shorten timelines without compromising scientific depth. In that environment, Syngene’s D2B platform is well-positioned. It offers the speed advantage of direct screening, the discipline of automated QC, and the biological depth needed to connect chemistry with real project decisions. Traditional hit-to-lead medicinal chemistry will remain essential, especially when purified compounds and detailed optimization are required. But for early-stage acceleration, Syngene’s D2B capability offers a sharper, more agile route from idea to evidence.
Ready to elevate your early discovery strategy? Syngene’ s DirecttoBiology approach unites design, microscale synthesis, automated analytics, and biology to streamline the discovery cycle. See how our integrated platform empowers smarter prioritization and accelerates the path from concept to candidate.
Learn More About Our Discovery Capabilities
Contributors: Tony Kurissery Anthappan, Gajjala Sravani, Muruganantham Rajendran, Santoshkumar N. Patil, Ardra P, Prafull Kumarsingh, Praveen Gopinathan Nair and Tripuraneni Srinivas
Contributors: Tony Kurissery Anthappan, Gajjala Sravani, Muruganantham Rajendran, Santoshkumar N. Patil, Ardra P, Prafull Kumarsingh, Praveen Gopinathan Nair and Tripuraneni Srinivas